Analysis¶
In this tutorial, we show the user how to instantiate the biceps.Analysis class which uses MBAR to get predicted populations of each conformational states and compute a BICePs score. We also provide a short description of the output data from analysis and embed the figures of posterior distribution of populations & nuisance parameters. Please refer to the documentation of
Analysis for more specific details.
[6]:
import biceps
import warnings
warnings.filterwarnings("ignore", category=UserWarning)
warnings.filterwarnings('ignore', category=FutureWarning)
[10]:
%matplotlib inline
A = biceps.Analysis(outdir="results", nstates=100, verbose=True)
fig = A.plot(plottype="hist") # plottype="step")
Loading results/traj_lambda0.00.npz ...
Loading results/traj_lambda1.00.npz ...
not all state sampled, these states [ 0 3 4 5 8 9 11 13 14 15 16 18 19 20 21 22 23 24 25 26 27 28 29 31
34 40 41 42 43 44 48 49 51 52 53 54 55 57 60 61 62 64 69 71 72 73 74 76
77 78 81 82 83 86 87 88 89 95 96 97 98 99] are not sampled
Loading results/traj_lambda0.00.pkl ...
Loading results/traj_lambda1.00.pkl ...
lam = [0.0, 1.0]
nstates 100
Time for MBAR: 0.077 s
Writing results/BS.dat...
Writing results/populations.dat...
Top 10 states: [46, 85, 92, 45, 39, 80, 65, 90, 59, 38]
Top 10 populations: [0.01001833 0.0212999 0.02792732 0.03482791 0.07746092 0.07924925
0.08826894 0.09715671 0.20440265 0.32912632]
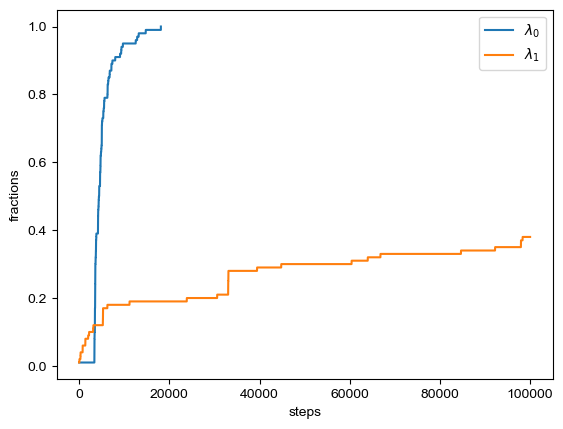
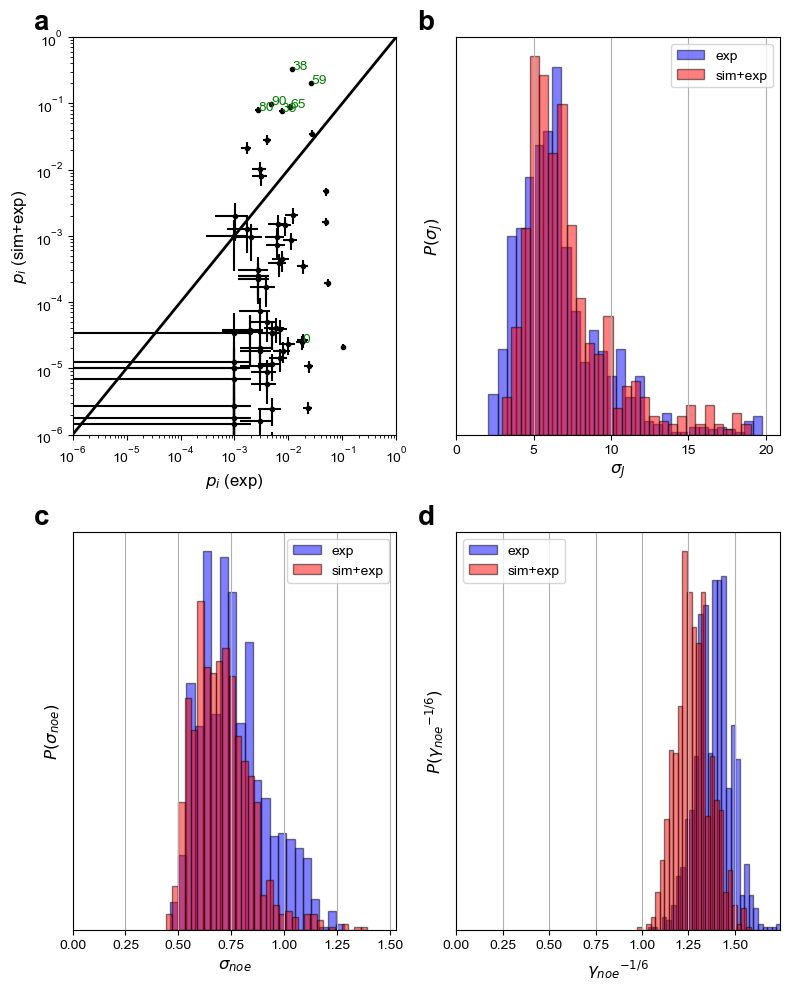
The output files include: population information (“populations.dat”), figure of sampled parameters distribution (“BICePs.pdf”), BICePs score information (“BS.dat”), which are shown above.
Now, let’s take a look at the populations file:
There are 100 rows corresponding to 100 clustered states. 4 columns corresponding to populations of each state (row) for 2 lambda values (first 2 columns) and population change (last 2 columns).
[11]:
import pandas as pd
import numpy as np
pops = np.loadtxt('results/populations.dat')
df = pd.DataFrame(pops)
df
[11]:
| 0 | 1 | 2 | 3 | |
|---|---|---|---|---|
| 0 | 0.017975 | 2.496320e-05 | 0.004195 | 5.931122e-06 |
| 1 | 0.009977 | 2.292886e-05 | 0.003138 | 7.283074e-06 |
| 2 | 0.002693 | 3.068112e-04 | 0.001553 | 1.773218e-04 |
| 3 | 0.000990 | 1.008111e-05 | 0.000989 | 1.008553e-05 |
| 4 | 0.000000 | 0.000000e+00 | NaN | NaN |
| ... | ... | ... | ... | ... |
| 95 | 0.006000 | 1.247277e-11 | 0.002441 | 5.105694e-12 |
| 96 | 0.000993 | 6.847524e-06 | 0.000993 | 6.850553e-06 |
| 97 | 0.004988 | 1.150700e-05 | 0.002225 | 5.157574e-06 |
| 98 | 0.004000 | 4.474545e-11 | 0.001996 | 2.241289e-11 |
| 99 | 0.002999 | 5.949074e-07 | 0.001729 | 3.439323e-07 |
100 rows × 4 columns
Conclusion
In this tutorial, the user learned how to call on the biceps.Analysis class in order to analyze the trajectory data, which automatically plots the posterior distribution of populations & nuisance parameters.
We now conclude the series of tutorials: Preparation, Restraint, PosteriorSampler and Analysis To learn more about BICePs, please check out our other examples & tutorials here.
# NOTE: The following cell is for pretty notebook rendering
[12]:
from IPython.core.display import HTML
def css_styling():
styles = open("../../../theme.css", "r").read()
return HTML(styles)
css_styling()
[12]: